The application of proteomics contributes to our understanding of the past, and it also brings us closer to ancient people and their lives—including murderous princes and their tear ducts.

The literary figure of Count Dracula is widely believed to be based on the Romanian Prince Vlad III. The famed medieval impaler was also known as Vlad Drăculea, which translates as Son of the Dragon, or simply devil. He is also attributed with a death count of more than 80,000 people (hence the "Vlad the Impaler" moniker), and he was rumored to shed tears of blood. These snippets of a real-life monster may be what inspired Bram Stoker to name his vampire character in the late 1890s.
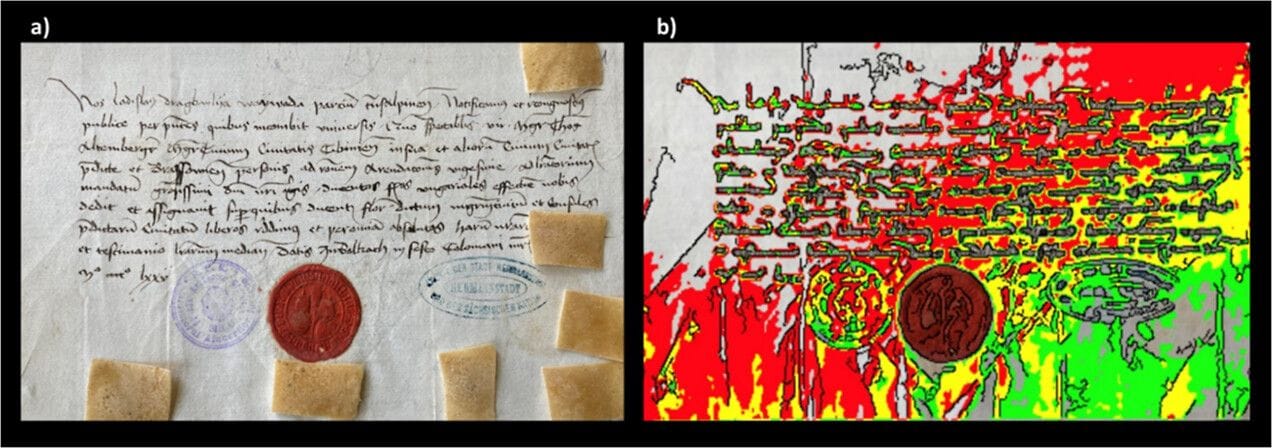
But beyond the mysterious and murderous lore, the prince himself left behind an array of tangible letters and documents. Such immortalization—in print, at least—garnered the attention of a proteomics-focused research team interested in ancient biomolecules on artifacts. The researchers used these letters, alongside new analytical techniques, to help "resurrect" the man and learn more about who he was.
Ancient biomolecules and proteins can open a window into the past, improving understanding of diverse fields such as evolutionary history, diet, and disease. For their study, the team used films of ethylene-vinyl acetate studded with strong cation and anion exchangers plus hydrophobic resins—a promising non-invasive technique which can extract proteins and small molecules from surfaces.

Count Dracula Resurrected: Proteomic Analysis of Vlad III the Impaler’s Documents by EVA Technology and Mass Spectrometry
DOI: 10.1021/acs.analchem.3c01461
After successfully harvesting peptides and proteins from three of Vlad Dracula's 15th century letters and characterizing them via mass spectrometry, the authors identified 575 human-related peptides, plus a further 2,212 from bacteria, viruses, fungi, insects, and plants. Of these, 165 peptides were classified as potential contaminants based on database information. In comparison, a modern control letter yielded 359 human peptides and 1,097 peptides across the other categories. Overall, in each individual category, only about 20 peptides were shared between the ancient and modern letters.
Next, the team focused on the proteins with the most advanced deamidation, since these degraded samples would be the oldest (and therefore most likely to be from Vlad). The results found 16 human proteins that were consistent across all three ancient letters. These reveal the prince could have suffered from respiratory and inflammatory issues, and possibly a rare condition called haemolacria. Excitingly, this would pinpoint the medical cause of his bloody tears described in the historical record.
Other proteins indicated he could have been exposed to Yersinia pestis, a plague bacteria, as well as viruses spread by ticks and mosquitoes. Although many people could have touched the letters—from those who opened and read them, to those who stored them in the archive—the authors are confident that the most prominent ancient proteins should be related to Prince Vlad the Impaler, who wrote and signed all three letters across a period of about 20 years.
The same team have looked at protein signatures in other ancient sources, albeit slightly less notorious ones. This includes cheese found in the tomb of Ptahmes in Egypt. This little snack for the afterlife had been wrapped in cloth and packed away in a jar more than 3,220 years ago. Proteomic analysis allowed the researchers to characterize about 500 peptides from over 90 proteins, most from human skin and saliva and likely due to contamination. The sample also yielded 9 peptides from cow, sheep, goat, or buffalo, including residues from albumin and lysozyme, which are found in milk.
This application of chemistry to archaeology was able to confidently suggest that the sample was a cheese-like product made using cow’s milk mixed with that from a goat or sheep. The analysis also revealed evidence of brucellosis—an infection caught from unpasteurized milk and cheese—which ties in with physical findings of the infection on mummies in other tombs.

Proteomic Analyses on an Ancient Egyptian Cheese and Biomolecular Evidence of Brucellosis
DOI: 10.1021/acs.analchem.8b02535
Explore More Proteomics Research from ACS Journals
An Introduction to Mass Spectrometry-Based Proteomics
Shuken, S. R. J. Proteome Res. 2023, 22, 7, 2151–2171
DOI: 10.1021/acs.jproteome.2c00838
TurboID-EV: Proteomic Mapping of Recipient Cellular Proteins Proximal to Small Extracellular Vesicles
Li, Y. et al. Anal. Chem. 2023, 95, 38, 14159–14164
DOI: 10.1021/acs.analchem.3c01015
Toward an Integrated Machine Learning Model of a Proteomics Experiment
Neely, B. A. et al. J. Proteome Res. 2023, 22, 3, 681–696
DOI: 10.1021/acs.jproteome.2c00711
Assessment and Prediction of Human Proteotypic Peptide Stability for Proteomics Quantification
Chiva, C. et al. Anal. Chem. 2023, 95, 37, 13746–13749
DOI: 10.1021/acs.analchem.3c02269
Assessing the Role of Trypsin in Quantitative Plasma and Single-Cell Proteomics toward Clinical Application
Woessmann, J. et al. Anal. Chem. 2023, 95, 36, 13649–13658
DOI: 10.1021/acs.analchem.3c02543
Putting Humpty Dumpty Back Together Again: What Does Protein Quantification Mean in Bottom-Up Proteomics?
Plubell, D. L. et al. J. Proteome Res. 2022, 21, 4, 891–898
DOI: 10.1021/acs.jproteome.1c00894
NMR-Based Methods for Protein Analysis
Hu, Y. et al. Anal. Chem. 2021, 93, 4, 1866–1879
DOI: 10.1021/acs.analchem.0c03830
