Unraveling the intricate composition of seawater has been an inherent challenge, but recent studies are opening up new ways to capture and identify a multitude of molecules in the oceans.

Seawater is vastly more complex than just being salty water, containing a multitude of molecules with diverse structures and origins that influence ecosystem functions. But analyzing its chemical composition has been challenging because of the inherent complexity and dynamic nature of marine environments. It can be hard to identify molecules in seawater due to their unknown or multiple biosynthetic origins and possible (bio)transformations, as well as the lack of available commercial standards or open-access spectral data. A further key challenge is that purification attempts are hindered by the low abundance in collected samples.
Therefore, techniques that enable capture of molecules as soon as they are released are critical to better understand origins and functional roles. To support this, the field of marine metabolomics is expanding—and previous research has shown that computational approaches such as genome and metabolome mining are becoming essential to researching natural products and molecules in the oceans.1
Now, work published in ACS Central Science showcases a handheld underwater solid-phase extraction instrument called I-SMEL, which can not only rapidly capture molecules within chemical seascapes but can also enrich the levels of specialized exometabolites by concentrating diluted molecules from large volumes of seawater.2
These exometabolites are important because they are released by keystone marine species such as sponges and can impact the chemical composition of their surroundings, and the same team have previously published on the role of exometabolites as new chemical reservoirs that could be collected from the water column while preserving marine biodiversity.3
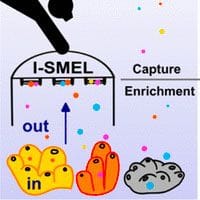
In Situ Capture and Real-Time Enrichment of Marine Chemical Diversity
DOI: 10.1021/acscentsci.3c00661
The new I-SMEL approach they have developed aims to sustainably access these under natural marine conditions, trapping the molecules of interest without harvesting the organisms and using a metabolomic approach based on mass spectometry to analyze the enriched extracts. The increased availability of high-resolution mass spectrometry in chemical analysis has dramatically improved the detection and identification of compounds in environmental samples.4
The team tested the instrument in a Mediterranean ecosystem with shaded parts dominated by sponges, with a particular focus on three common species. Results showed the I-SMEL device was able to produce a chemical fingerprint, highlighting diversity and proportions of exometabolites in the samples. But the real advantage here is that I-SMEL is a handheld tool that can be easily used at various depths by scientific divers.
These kinds of techniques will help ecologists in identifying key molecules used for communication between organisms such as corals and algae as well as sponges, and they will further support arguments for the preservation of marine ecosystems. Even so, more discussion is needed across disciplines to agree how to identify tentative or unknown compounds via literature and databases for the benefit of all investigations.
References
- Kim, H. W. et al. NPClassifier: A Deep Neural Network-Based Structural Classification Tool for Natural Products. J. Nat. Prod. 2021, 84, 11, 2795–2807.
- Mauduit, M. et al. In Situ Capture and Real-Time Enrichment of Marine Chemical Diversity. ACS Cent. Sci. 2023, 9, 11, 2084–2095.
- Mauduit, M. et al. Diving into the Molecular Diversity of Aplysina cavernicola’s Exometabolites: Contribution of Bromo-Spiroisoxazoline Alkaloids. ACS Omega 2022, 7, 47, 43068–43083.
- Schymansk, E. L. et al. Identifying Small Molecules via High Resolution Mass Spectrometry: Communicating Confidence. Environ. Sci. Technol. 2014, 48, 4, 2097–2098.
